Simulate Data
1 Preamble
1.1 Load Libraries
2 Simulate Data
These data were simulated for the workshop with the help of AI.
Code
set.seed(52242)
# -----------------------------
# 1. Settings
# -----------------------------
N <- 250
timepoints <- 1:4
ages <- 14:17
n_time <- length(ages)
# Participant-level info
id <- 1:N
sexMale <- rbinom(N, 1, 0.5) # 0=female, 1=male
df <- expand.grid(ID = id, timepoint = timepoints) |>
arrange(ID, timepoint) |>
mutate(sexMale = rep(sexMale, each = n_time))
df$age <- NA
df$age[which(df$timepoint == 1)] <- 14
df$age[which(df$timepoint == 2)] <- 15
df$age[which(df$timepoint == 3)] <- 16
df$age[which(df$timepoint == 4)] <- 17
# -----------------------------
# 2. Random intercepts (person-level)
# -----------------------------
# Correlated latent traits: depression, anxiety, sleep, SES
Sigma_u <- matrix(c(
6.0, 3.0, -0.5, -0.3,
3.0, 6.0, -0.4, -0.3,
-0.5,-0.4, 1.0, 0.5,
-0.3,-0.3, 0.5, 1.0
), nrow=4, byrow=TRUE)
u <- mvrnorm(N, mu=c(0,0,0,0), Sigma=Sigma_u)
colnames(u) <- c("u_dep","u_anx","u_sleep","u_ses")
df <- df |>
left_join(data.frame(ID=id, u_dep=u[,1], u_anx=u[,2], u_sleep=u[,3], u_ses=u[,4]), by="ID")
# -----------------------------
# 3. Time-varying residuals
# -----------------------------
Sigma_e <- matrix(c(
2.0, 1.0, -0.3, -0.2,
1.0, 2.0, -0.2, -0.2,
-0.3,-0.2, 0.5, 0.3,
-0.2,-0.2, 0.3, 0.5
), nrow=4, byrow=TRUE)
res <- lapply(1:N, function(i){
mvrnorm(n_time, mu=c(0,0,0,0), Sigma=Sigma_e)
})
res <- do.call(rbind, res)
colnames(res) <- c("e_dep","e_anx","e_sleep","e_ses")
df <- bind_cols(df, as.data.frame(res))
# -----------------------------
# 4. Fixed effects
# -----------------------------
beta_age_dep <- 1.0
beta_age_anx <- 0.8
beta_sex_dep <- -2
beta_sex_anx <- -2
beta_age_sex_dep <- -0.4
beta_age_sex_anx <- -0.3
beta_ses_dep <- -0.5
beta_ses_anx <- -0.5
beta_sleep_dep <- -0.3
beta_sleep_anx <- -0.2
# -----------------------------
# 5. Compute SES (z-scored)
# -----------------------------
df$ses_cont <- df$u_ses + df$e_ses
df$ses <- as.numeric(scale(df$ses_cont))
# -----------------------------
# 6. Compute Sleep (5-10 hrs)
# -----------------------------
df$sleep_cont <- df$u_sleep + df$e_sleep
df$sleep <- pmin(pmax(8 + df$sleep_cont, 5), 10)
# -----------------------------
# 7. Latent depression & anxiety with cross-lagged structure
# -----------------------------
# Sort correctly
df <- df |> arrange(ID, age)
# Parameters for temporal dynamics
phi_dep <- 0.50 # autoregressive depression
phi_anx <- 0.50 # autoregressive anxiety
cl_anx_to_dep <- 0.35 # anxiety -> later depression (stronger)
cl_dep_to_anx <- 0.10 # depression -> later anxiety (weaker)
# Initialize latent variables
df$depression_latent <- NA
df$anxiety_latent <- NA
for (i in unique(df$ID)) {
idx <- which(df$ID == i)
for (t in seq_along(idx)) {
row <- idx[t]
# Baseline (age 14)
if (t == 1) {
df$depression_latent[row] <-
df$u_dep[row] +
beta_sex_dep * df$sexMale[row] +
beta_ses_dep * df$ses[row] +
beta_sleep_dep * df$sleep[row] +
df$e_dep[row]
df$anxiety_latent[row] <-
df$u_anx[row] +
beta_sex_anx * df$sexMale[row] +
beta_ses_anx * df$ses[row] +
beta_sleep_anx * df$sleep[row] +
df$e_anx[row]
} else {
prev <- idx[t - 1]
df$depression_latent[row] <-
df$u_dep[row] +
phi_dep * df$depression_latent[prev] +
cl_anx_to_dep * df$anxiety_latent[prev] +
(beta_age_dep + beta_age_sex_dep * df$sexMale[row]) * (df$age[row] - 14) +
beta_sex_dep * df$sexMale[row] +
beta_ses_dep * df$ses[row] +
beta_sleep_dep * df$sleep[row] +
df$e_dep[row]
df$anxiety_latent[row] <-
df$u_anx[row] +
phi_anx * df$anxiety_latent[prev] +
cl_dep_to_anx * df$depression_latent[prev] +
(beta_age_anx + beta_age_sex_anx * df$sexMale[row]) * (df$age[row] - 14) +
beta_sex_anx * df$sexMale[row] +
beta_ses_anx * df$ses[row] +
beta_sleep_anx * df$sleep[row] +
df$e_anx[row]
}
}
}
# -----------------------------
# 8. Transform to skewed counts (preserve ordering & skew)
# -----------------------------
df <- df |>
mutate(
dep_rank = rank(depression_latent, ties.method = "average") / (n() + 1),
anx_rank = rank(anxiety_latent, ties.method = "average") / (n() + 1),
depression = round(pmin(qgamma(dep_rank, shape = 2, scale = 5), 30)),
anxiety = round(pmin(qgamma(anx_rank, shape = 2, scale = 5), 30))
) |>
select(-dep_rank, -anx_rank)
# -----------------------------
# 9. Introduce attrition
# -----------------------------
df <- df |>
mutate(
p_dropout = case_when(
age == 14 ~ 0.00,
age == 15 ~ 0.05,
age == 16 ~ 0.10,
age == 17 ~ 0.15
),
dropout = rbinom(n(), 1, p_dropout)
) |>
mutate(
ses = ifelse(dropout == 1, NA, ses),
sleep = ifelse(dropout == 1, NA, sleep),
depression = ifelse(dropout == 1, NA, depression),
anxiety = ifelse(dropout == 1, NA, anxiety)
) |>
select(-p_dropout, -dropout)
# -----------------------------
# 10. Factorize sex for plotting
# -----------------------------
df$sex <- NA
df$sex[which(df$sexMale == 0)] <- "female"
df$sex[which(df$sexMale == 1)] <- "male"
df$sexEffectCode <- NA
df$sexEffectCode[which(df$sexMale == 0)] <- -1 # female
df$sexEffectCode[which(df$sexMale == 1)] <- 1 # male
# -----------------------------
# 11. Final dataset
# -----------------------------
df <- df |>
select(ID, timepoint, age, sex, sexMale, sexEffectCode, ses, sleep, depression, anxiety)2.1 Check
Code
describe(df) depression anxiety sleep ses
depression 1.0000000 0.7352566 -0.3022551 -0.3171477
anxiety 0.7352566 1.0000000 -0.2471106 -0.2497336
sleep -0.3022551 -0.2471106 1.0000000 0.4948693
ses -0.3171477 -0.2497336 0.4948693 1.0000000Code
head(df)Linear mixed model fit by REML. t-tests use Satterthwaite's method [
lmerModLmerTest]
Formula: depression ~ lag(anxiety) + lag(depression) + age + sex + (1 |
ID)
Data: df
REML criterion at convergence: 4775.6
Scaled residuals:
Min 1Q Median 3Q Max
-3.2002 -0.5081 -0.1284 0.4673 3.3306
Random effects:
Groups Name Variance Std.Dev.
ID (Intercept) 21.83 4.672
Residual 7.16 2.676
Number of obs: 866, groups: ID, 250
Fixed effects:
Estimate Std. Error df t value Pr(>|t|)
(Intercept) 1.01595 1.44137 715.47955 0.705 0.48113
lag(anxiety) 0.10589 0.02851 708.01306 3.714 0.00022 ***
lag(depression) 0.18044 0.02891 717.42565 6.241 7.43e-10 ***
age 0.49832 0.08698 605.87838 5.729 1.59e-08 ***
sexmale -3.82035 0.62760 229.13169 -6.087 4.79e-09 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Correlation of Fixed Effects:
(Intr) lg(nx) lg(dp) age
lag(anxity) -0.045
lag(dprssn) -0.120 -0.766
age -0.946 -0.001 0.067
sexmale -0.211 0.015 0.042 0.009Linear mixed model fit by REML. t-tests use Satterthwaite's method [
lmerModLmerTest]
Formula: anxiety ~ lag(depression) + lag(anxiety) + age + sex + (1 | ID)
Data: df
REML criterion at convergence: 4812.4
Scaled residuals:
Min 1Q Median 3Q Max
-3.2445 -0.5619 -0.1192 0.4608 3.4108
Random effects:
Groups Name Variance Std.Dev.
ID (Intercept) 20.639 4.543
Residual 7.736 2.781
Number of obs: 866, groups: ID, 250
Fixed effects:
Estimate Std. Error df t value Pr(>|t|)
(Intercept) -3.53258 1.48987 707.97997 -2.371 0.018 *
lag(depression) 0.04526 0.02987 732.31955 1.516 0.130
lag(anxiety) 0.21592 0.02947 722.36199 7.328 6.27e-13 ***
age 0.81105 0.09034 609.49541 8.978 < 2e-16 ***
sexmale -3.91681 0.61448 231.11897 -6.374 9.87e-10 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Correlation of Fixed Effects:
(Intr) lg(dp) lg(nx) age
lag(dprssn) -0.120
lag(anxity) -0.046 -0.765
age -0.950 0.066 -0.001
sexmale -0.202 0.045 0.016 0.010Code
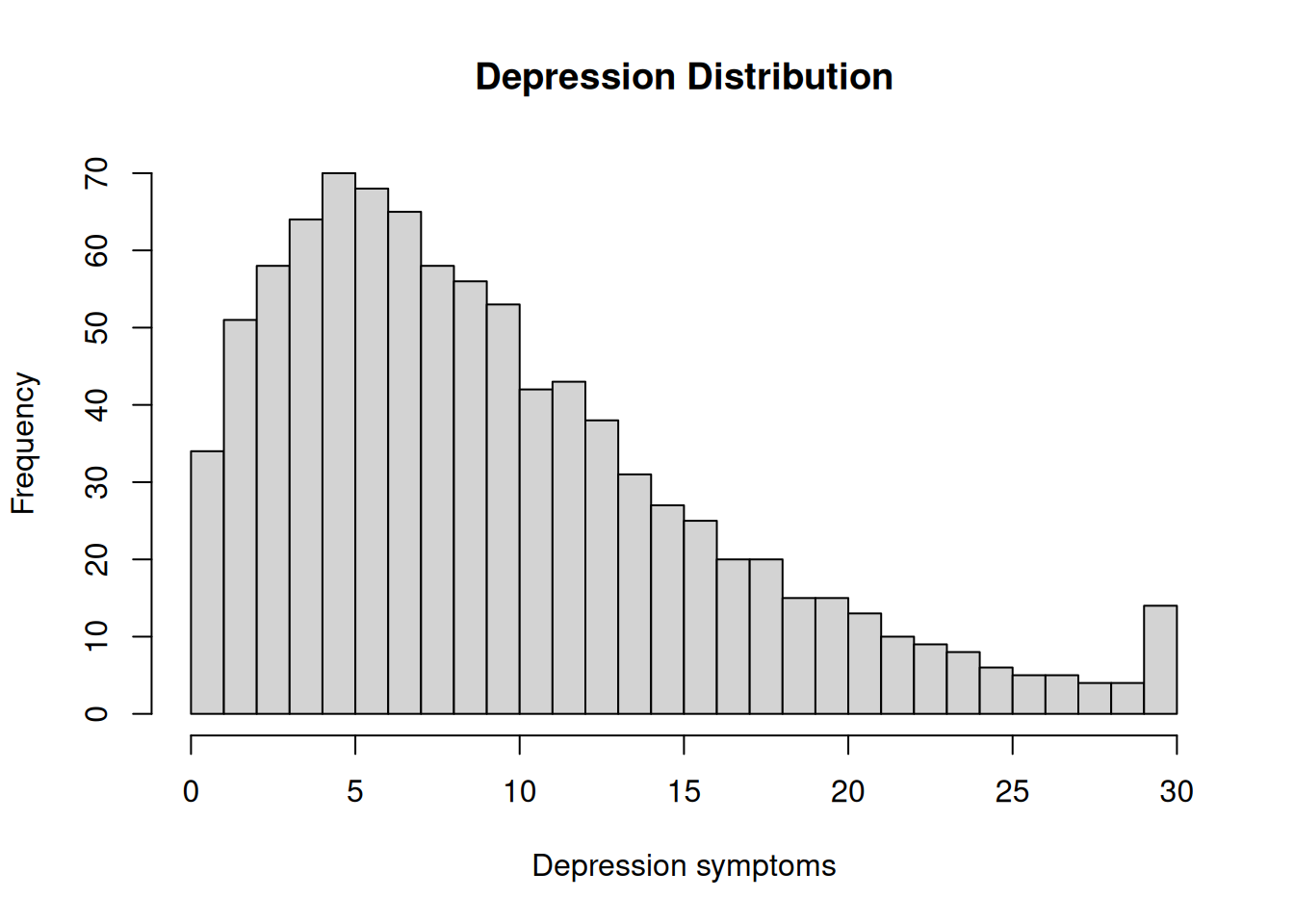
hist(
df$depression,
breaks = 30,
main = "Depression Distribution",
xlab = "Depression symptoms")Code
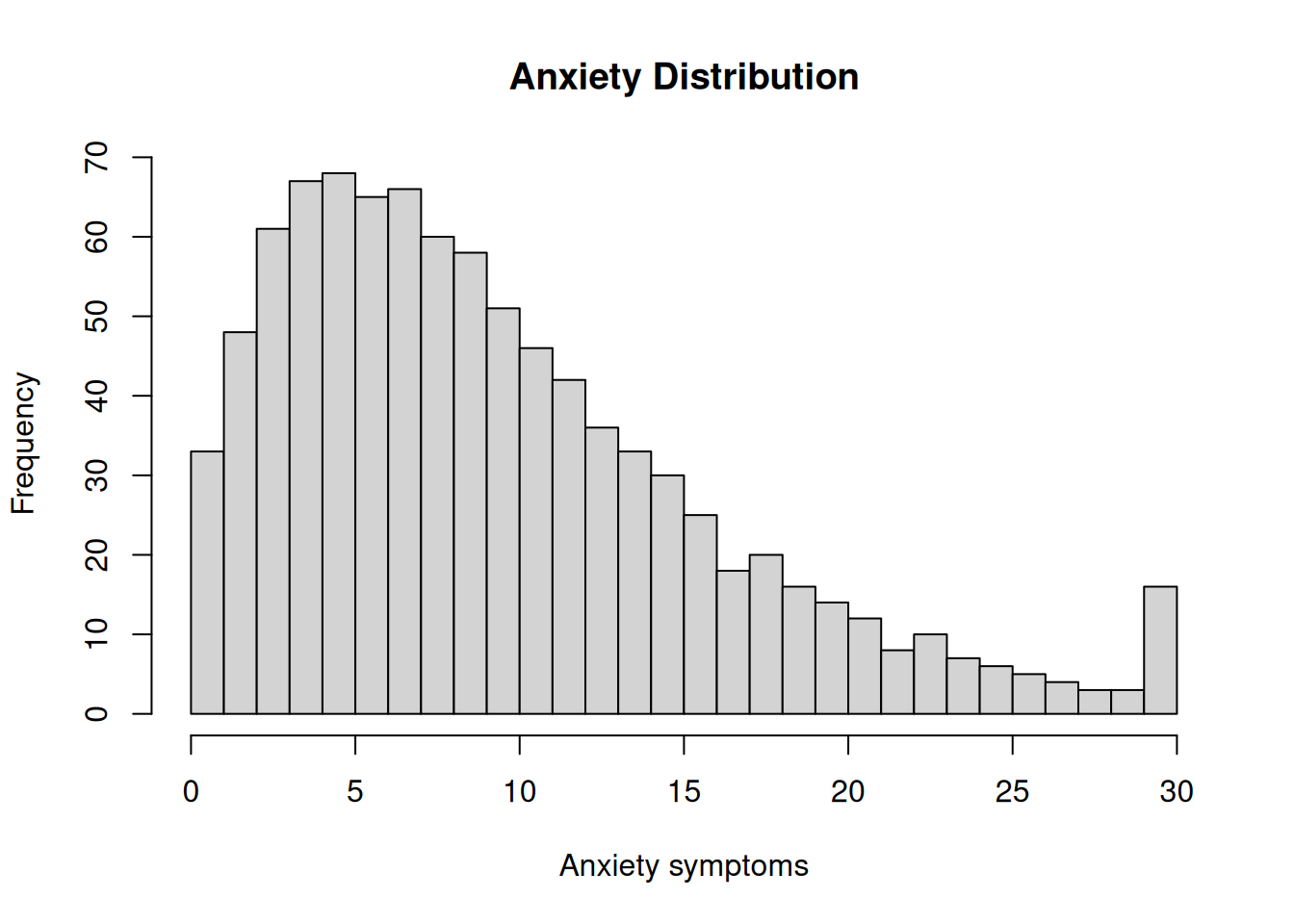
hist(
df$anxiety,
breaks = 30,
main = "Anxiety Distribution",
xlab = "Anxiety symptoms")Code
# --- Summarize mean + SD by age and sex ---
df_summary <- df |>
group_by(age, sex) |>
summarize(
mean_dep = mean(depression, na.rm=TRUE),
sd_dep = sd(depression, na.rm=TRUE),
mean_anx = mean(anxiety, na.rm=TRUE),
sd_anx = sd(anxiety, na.rm=TRUE),
n = n(),
.groups = "drop"
)
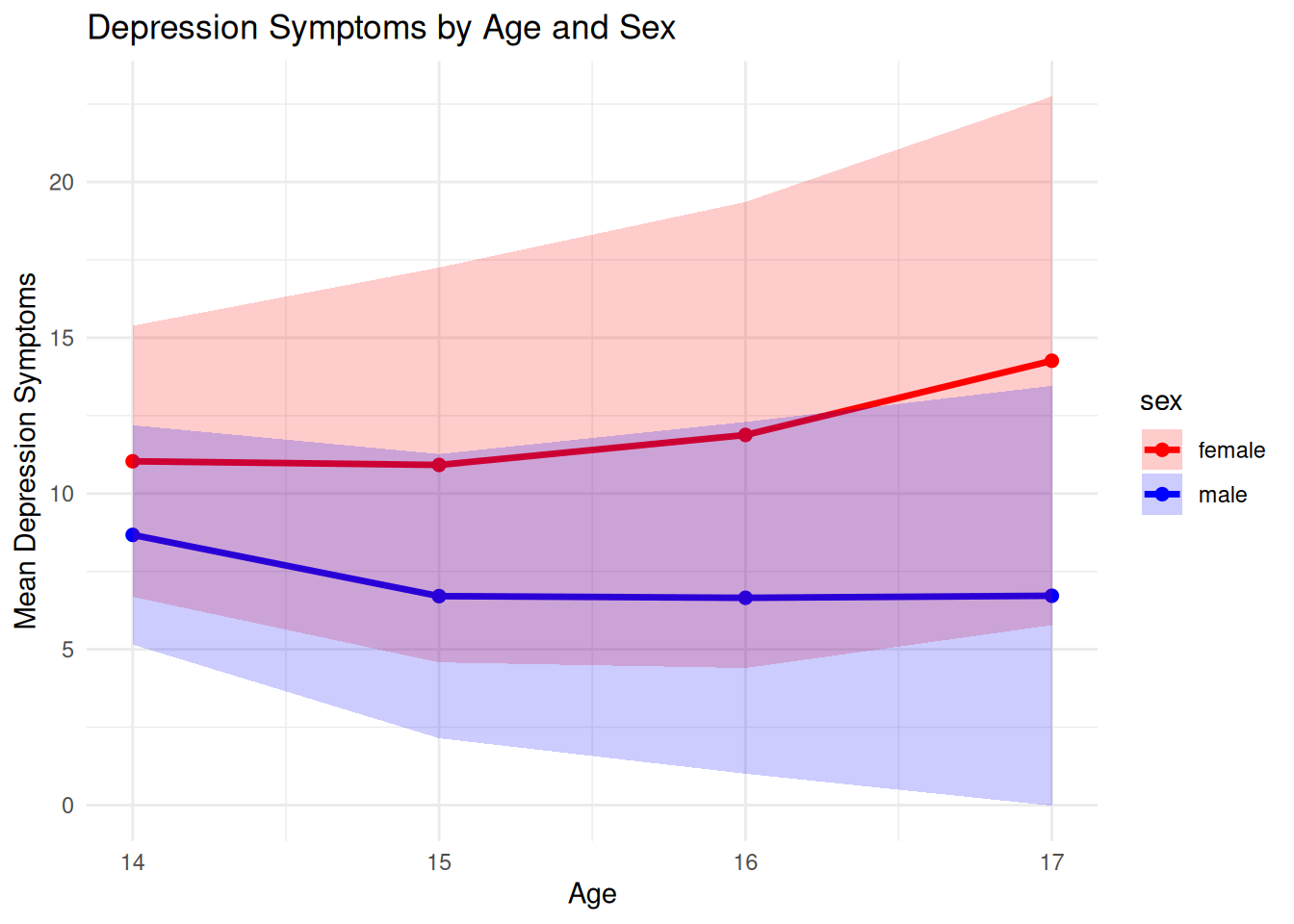
# --- Plot Depression by Age and Sex ---
ggplot(df_summary, aes(x=age, y=mean_dep, color=sex)) +
geom_line(linewidth=1.2) +
geom_point(size=2) +
geom_ribbon(aes(ymin=mean_dep-sd_dep, ymax=mean_dep+sd_dep, fill=sex), alpha=0.2, color=NA) +
labs(title="Depression Symptoms by Age and Sex",
x="Age", y="Mean Depression Symptoms") +
theme_minimal() +
scale_color_manual(values=c("female"="red", "male"="blue")) +
scale_fill_manual(values=c("female"="red", "male"="blue"))Code
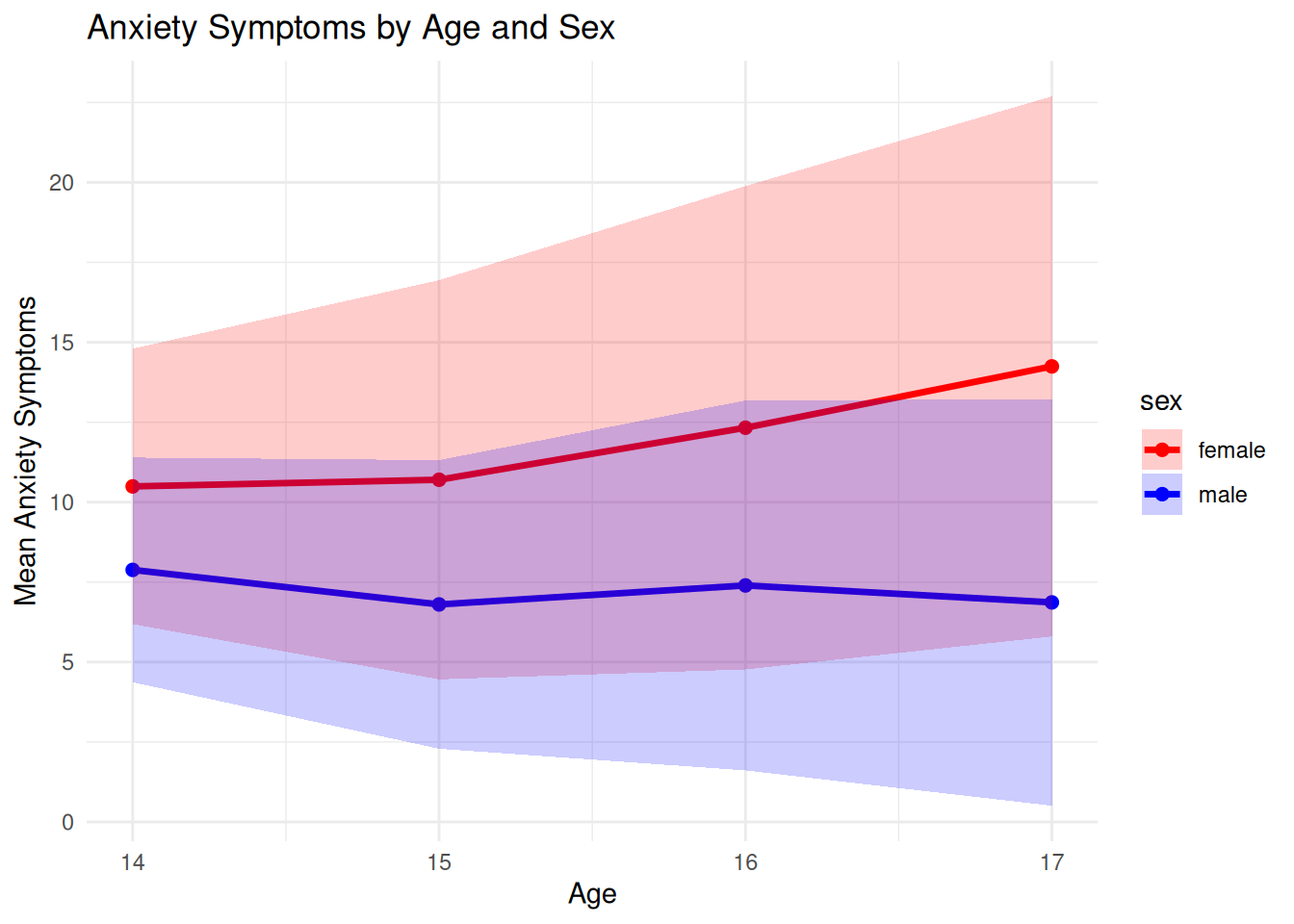
# --- Plot Anxiety by Age and Sex ---
ggplot(df_summary, aes(x=age, y=mean_anx, color=sex)) +
geom_line(linewidth=1.2) +
geom_point(size=2) +
geom_ribbon(aes(ymin=mean_anx-sd_anx, ymax=mean_anx+sd_anx, fill=sex), alpha=0.2, color=NA) +
labs(title="Anxiety Symptoms by Age and Sex",
x="Age", y="Mean Anxiety Symptoms") +
theme_minimal() +
scale_color_manual(values=c("female"="red", "male"="blue")) +
scale_fill_manual(values=c("female"="red", "male"="blue"))3 Save Data
Code
write.csv(
x = df,
file = "./data/mydata_long.csv",
row.names = FALSE)